Biology of small populations with Drop-Based Microfluidics
Biological populations consist of many distinct members. Characterizing this diversity is critical for understanding and perturbing phenomena such as evolution, organismal development, cancer and immune response. This requires identification of the key individuals that drive these phenomena and the specific genes or functions that control their behavior. This can only be achieved with individual access to each member of the population. However, most biological assays sample a large number of cells simultaneously; thus, individual properties of unique cells cannot be discerned from these averages. My goal is to develop methods to efficiently assay large numbers of single cells or microorganisms and to apply these methods to understand important biological processes that critically depend on the variation between individuals.
Measuring individuals – be they microorganisms, cells or molecules – requires different tools from those used to assay whole populations. When a single cell is sampled in the same volume as that used to sample a millions cells, the effective concentration of its content is a million times smaller, while the concentration of contaminants or other inhibiting agents remains the same. Drop-Based Microfluidics (DBM) is used to perform single-cell assays by encapsulating each cell in a micron sized drop of water immersed in an inert carrier oil. To perform the assay, drops can be split, merged, refilled, incubated and thermocycled at rates of thousands per second with exquisite control over individual drops. To measure the results, the fluorescence intensity of each drop can be detected and analyzed using a custom setup; moreover drops can also be sorted in a fashion similar to Fluorescently Activated Cell Sorters (FACS).
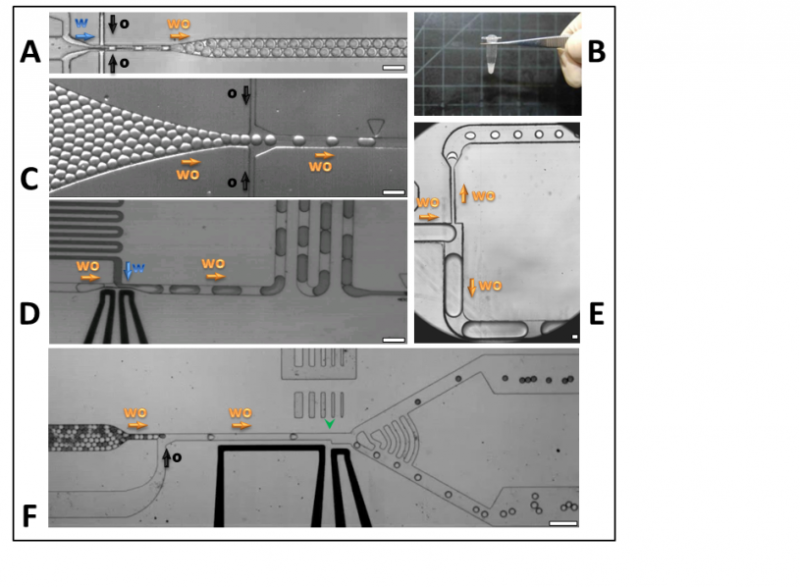
Figure 1 – Drop-Based Microfluidics. A) An aqueous solution is co-flowed with carrier oil to produce water-in-oil emulsion drops that encapsulate any cells, particles, or molecules present in the aqueous solution. B) One million drops, each 25 μm in diameter, are visible as ≈20 μL of emulsion stored in a 200 μL microtube. C) Previously formed drops are reinjected into a microfluidic device. D) Drops flow through a T-junction where a blue-dyed aqueous solution is injected into them by locally destabilizing the interface of the drops via electrodes (thick black lines) that produce an electric field. E) Drops flowing through a T-junction split into two drops with sizes determined by the geometry of the device. F) Drops are interrogated via a laser spot (below green arrowhead) and sorted via electrodes (thick black lines) according to the emitted fluorescence. All scale bars are 100 μm. Arrows indicate the direction of flow for different phases: aqueous fluid (w), carrier oil (o), and water-in-oil emulsion drops (wo). Figure taken from Guo et al. Lab on Chip 2012.
Evolution of small viral populations.
This work is performed with Huidan Zhang, Connie Chang, Ye Tao and in collaboration with Andrew Feldman, Shakhnovich lab, Pipas lab, Wobus lab, Smith lab and Kellogg Schwab.
A human viral disease can emerge when a single bird flu virus is able to infect a human host; characterizing the propagation of viral lineages is difficult but crucial for understanding, predicting and preventing the outburst of viral epidemics. A standard approach to probe this genetic propagation is by mutation accumulation experiments, in which a single lineage is propagated by plating a population of viruses, randomly picking a single colony to propagate, and repeating this process over time. Unfortunately, it is difficult to extract from these experiments enough information for a reliable characterization of an evolving species due to the small number of replicates collected and to the insufficient duration of the experiments. Using microfluidics we can perform millions of parallel mutation accumulation experiments in drops.
To accomplish this, I developed an “evolution chip”, which passes millions of populations per hour from one drop where they replicated in the previous generation to a new drop containing fresh nutrients and hosts, where they will replicate in the next generation. When we co-encapsulated viruses and host cells in the same drop, viruses readily replicate in drops and smaller populations yield a broader clonal distribution. After propagating lineages of small populations in a new environment for 4 generations we find a much broader distribution of adapted clones compared to conventional passaging of large populations as shown in Figure 2.
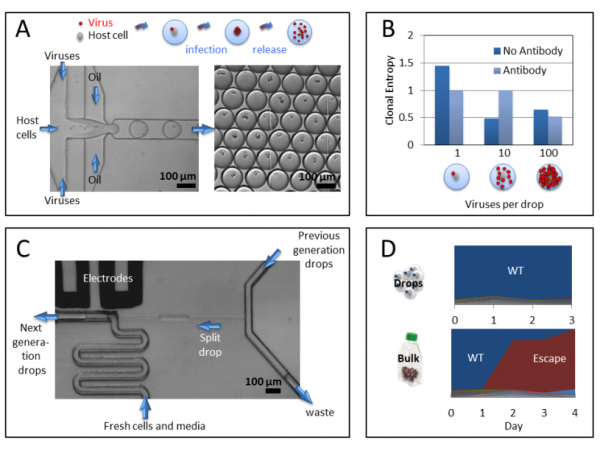
Figure 2 – Viral evolution in drops. A) Viruses and their host cells are co-encapsulated in drops, where they infect and replicate. B) The diversity of viral genomes obtained from millions of drops that initially contained only one virus is higher than that of drops that started with more than one virus, as expected in population bottlenecks. C) To propagate viral lineages in drops, a fraction of every drop from the previous generation is merged with the next generation drop containing fresh cells. D) In the presence of a drug, a single mutant escapes and takes over the population in bulk. In drops, many mutants evolve side by side and providing multiple examples of escape variants from a single experiment.
Evolution chip
High throughput Single Cell Labeling (Hi-SCL)
This work is a collaboration with Oren Ram and Noam Shoresh at the Bernstein lab.
Cells of identical genetic origin develop to perform a variety of different functions in our body. The proteins encoded in the genome are not all expressed at the same level in each cell, leading to this cell-to-cell variation. Characterizing the cellular modifications that lead to this variation is key to understanding cellular differentiation in our body. However, the measurements for such modifications are currently performed over a population of cells and it is impossible to use them to study variations between cells in the same population. To perform these assays at the resolution of a single-cell we developed a new technique, called High-throughput Single Cell Labeling (Hi-SCL), where each cell is uniquely labeled in a drop. After labeling, drops are pooled and the emulsion is broken to make a single sample, which undergoes an epigenomic assay and deep-sequencing. Upon sequencing, the labels attached to the cellular fragments are used to associate each fragment with the cell of origin, enabling deconvolution of the epigenomic data to 100 profiles of single cells. These profiles are sufficient to identify two cell types that were pre-mixed in the sample, as described in Figure 3. Thus, Hi-SCL allows a sample of many cells to be assayed simultaneously, yet analyzed at single-cell resolution.
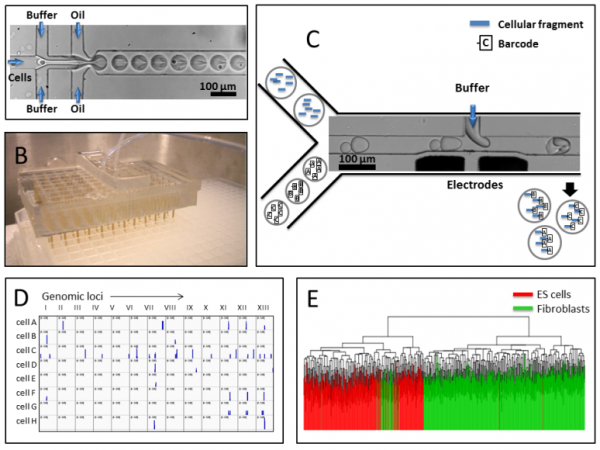
Figure 3 – High throughput Single-Cell Labeling. A) Using a microfluidic drop maker cells are encapsulated and lysed in drops. B) To label cells, barcodes are encapsulated in a parallel drop-maker that adapts into a quarter of a 384 well-plate. C) Every drop containing cell extract is merged with a drop containing high concentration of a unique barcode and a ligation buffer. D) Raw single-cell data: the occurrence of chromatin marks in 13 different genomic locations is plotted for 8 representative single cells. E). A thousand cells of two known types are separated based on their single-cell epigenome using a clustering algorithm.
Cell Encapsulation
3 Point Merger
Fabrication of multilayered microfluidic devices and its application for double emulsions
Photolithography is an accurate, reproducible and easy method for fabricating micron scale devices. The basic outcome of Photolithography is a single layer, and it is possible to repeat the exposure process and end up with multilayered landscapes. However, some topologies are impossible to achieve using multiple exposures and require a complementary method of stacking up devices after fabrication. We use a simple method for aligning stacks of micron-scale devices that relies on matching locks and keys that are an inherent part of the device. Applications of multilayered devices are numerous. We focus here on the generation of double emulsions that are useful for instance in encapsulation (drug delivery) and particle synthesis. The formation of double emulsions in single layered microfluidic devices demands precise spatial control over surface properties of the device channels. However in the case of axial symmetric microfluidic devices such as capillaries, this is no longer a constraint since there is no direct contact between the double emulsion and the device surface. We show that forming double emulsions in multilayered microfluidic device is similar to their formation in capillaries, alleviating the need for precise spatial control of wettability. Improving the robustness of double emulsion formation in photolithographic devices is important for large scale uses of double emulsions.
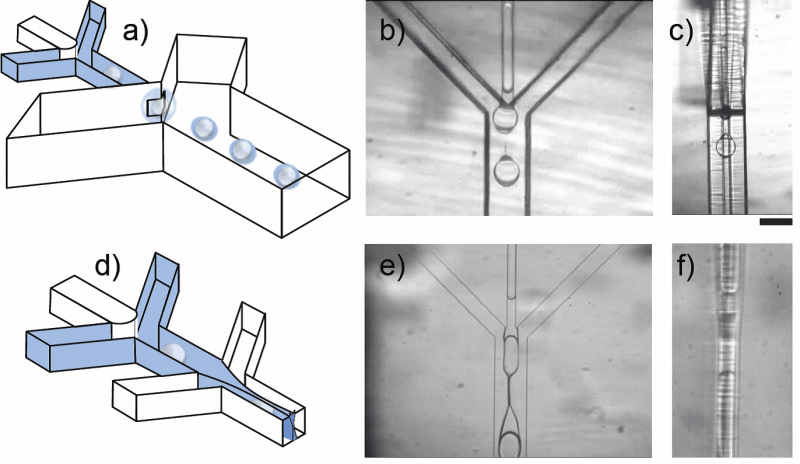
Figure 4 – Non-planar devices for making double emulsions. a) A schematic drawing of a hydrophobic non-planar device with water drops (white) forming at the first, planar junction and oil drops (blue) wrapping the water drops in the second, non-planar junction. b) Top view of a real non-planar junction, similar to the second junction in the schematics, as it is making drops. c) Side view of the same junction in (b). d) A schematic drawing of a hydrophobic planar device with water drops forming at the first junction and a sheath of oil that wets the floor and ceiling wrapping the water drops in the second junction. e) Top view of a real planar junction similar to the second junction in the schematics, as it is making drops. f) Side view of the same junction in (e). Scale bar is 100µm. Figure taken from Rotem et al. Lab on Chip 2012.
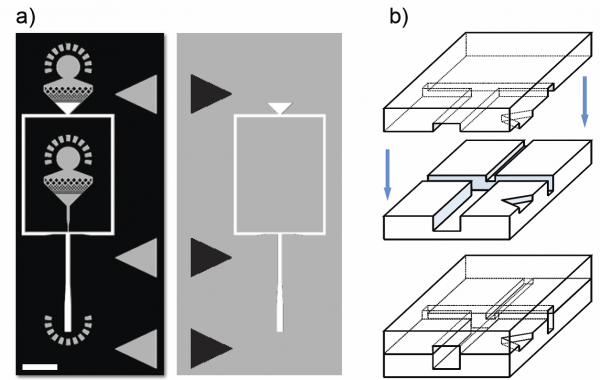
Figure 5 – Fabricating non-planar devices. a) Two photolithographic masters that are used for making the device. Each master has two layers: the base of the master is black, the first layer is gray and the second layer is white. Scale bar is 1mm. b) Two PDMS molds, the top one replicated from the right master in (a) and the bottom one replicated from the left master in (a) are bonded face to face, with the triangles on both sides matching to align the molds and make the final device. Scale bar is 100µm. Figure taken from Rotem et al. Lab on Chip 2012.
